October 16, 2019
Enzyme trigger that tells cells to move characterized for first time
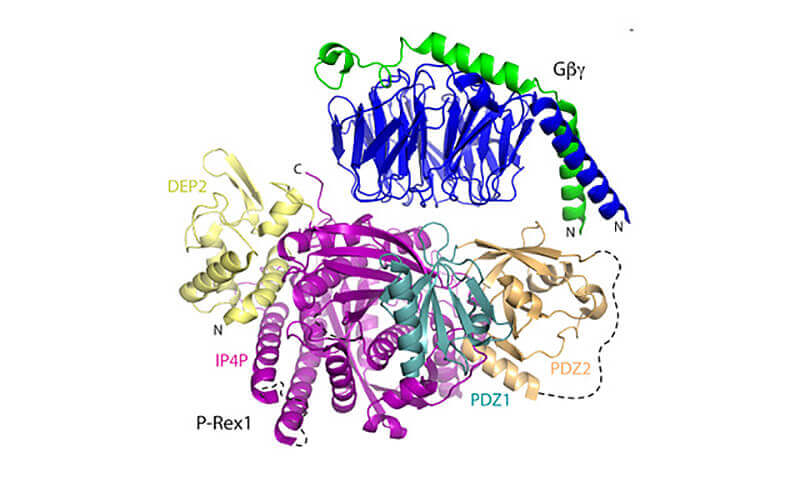 The physical structure of the enzyme that signals cells to move within the body—to fight infection or to spread cancer — has been identified by a Purdue-led team. The information is expected to be used to develop new drug therapies to prevent certain cancers from metastasizing. (Purdue University image/John Tesmer)
The physical structure of the enzyme that signals cells to move within the body—to fight infection or to spread cancer — has been identified by a Purdue-led team. The information is expected to be used to develop new drug therapies to prevent certain cancers from metastasizing. (Purdue University image/John Tesmer)
WEST LAFAYETTE, Ind. — Some cells have the ability to travel in the human body, and depending on the circumstances, this can be either very, very good or very, very bad.
White blood cells speeding toward an infection are the good guys in this story, while cancer cells spreading to metastasize and conquer wear the black hats.
The ability to control this movement could lead to drugs that could boost a body's response to an infection, or thwart the movement of cancer cells.
John J.G. Tesmer, Purdue University's Walther Professor in Cancer Structural Biology, is part of a research team that used an advanced electron microscope technique, cryo-electron microscopy, to determine the precise shape of the proteins involved. The research is published this week in Science Advances.
"Our structural data will allow the development of models for these proteins and their interactions, which could eventually lead to new therapies," Tesmer said.
The enzyme PREX1 allows cells to reorganize their cytoskeletons for movement within the body.
"During metastasis, many common cancers find a way to express the same enzyme which, in turn, helps them move around and colonize different niches in the body," Tesmer said. "By interfering with the enzyme function, we might have an anti-metastatic cancer strategy.
"But this is hard to do without knowledge of the structure of the enzyme or of how it is regulated by other molecules. This is why we are excited about these results."
The research team also was able to observe the enzyme bound to a protein that regulates it, known as G-betagamma.
"Moreover, the structure of PREX1 revealed it has a configuration not previously seen in eukaryotic organisms," Tesmer said. "It has unique properties that may allow us to selectively target pockets on its surface with designed chemical compounds that may block G-betagamma from binding."
The work aligns with Purdue's Giant Leaps celebration, acknowledging the university’s global advancements made in health, longevity and quality of life as part of Purdue’s 150th anniversary. This is one of the four themes of the yearlong celebration’s Ideas Festival, designed to showcase Purdue as an intellectual center solving real-world issues.
This work was funded by National Institutes of Health grants CA221289,
HL122416 and HL071818. Tesmer also received support from the Walther Cancer Foundation.
Writer: Steve Tally, 765-494-9809, steve@purdue.edu, @sciencewriter
Source: John J.G. Tesmer, 765-494-1807, jtesmer@purdue.edu, @tesmer_john
ABSTRACT
Cryo–electron microscopy structure and analysis of the P-Rex1–Gbg signaling scaffold
Jennifer N. Cash1, Sarah Urata2, Sheng Li2, Sandeep K. Ravala3, Larisa V. Avramova3, Michael D. Shost1, J. Silvio Gutkind4, John J. G. Tesmer3*, Michael A. Cianfrocco1*
PIP3-dependent Rac exchanger 1 (P-Rex1) is activated downstream of G protein–coupled receptors to promote neutrophil migration and metastasis. The structure of more than half of the enzyme and its regulatory G protein binding site are unknown. Our 3.2 Å cryo-EM structure of the P-Rex1–Gbg complex reveals that the carboxyl-terminal half of P-Rex1 adopts a complex fold most similar to those of Legionella phosphoinositide phosphatases. Although catalytically inert, the domain coalesces with a DEP domain and two PDZ domains to form an extensive docking site for Gbg. Hydrogen-deuterium exchange mass spectrometry suggests that Gbg binding induces allosteric changes in P-Rex1, but functional assays indicate that membrane localization is also required for full activation. Thus, a multidomain assembly is key to the regulation of P-Rex1 by Gbg and the formation of a membrane-localized scaffold optimized for recruitment of other signaling proteins such as PKA and PTEN.

