November 6, 2017
Research Highlights
Findings probe cell cooperation, ‘en masse’ migration
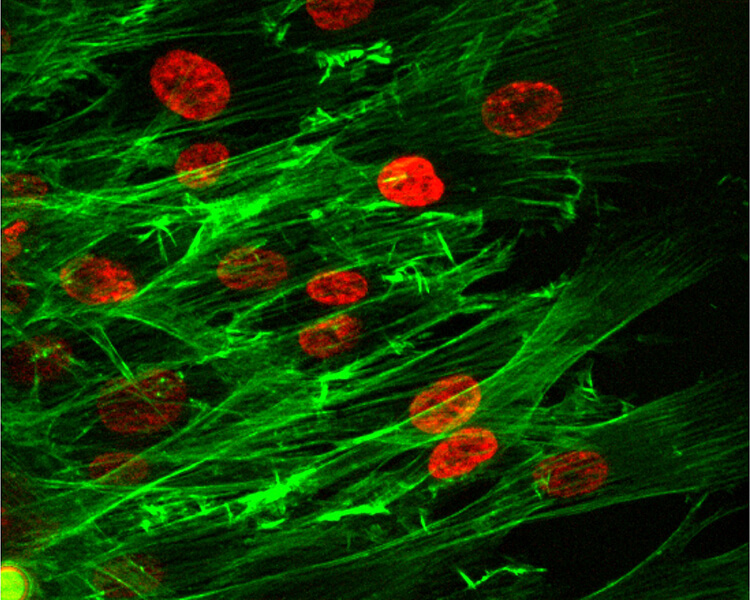 This image shows the group migration of cells, with red indicating the cell nuclei and the green representing the cell’s “cytoskeleton.” (Purdue University photo/Bumsoo Han)
Download image
This image shows the group migration of cells, with red indicating the cell nuclei and the green representing the cell’s “cytoskeleton.” (Purdue University photo/Bumsoo Han)
Download image
WEST LAFAYETTE, Ind. – New research findings are revealing secrets about how living cells “cooperate” with each other, joining into groups that migrate collectively and alter tissue.
The research focused on cells called fibroblasts, which are found in connective tissue and produce the “extracellular matrix,” a scaffold-like material between cells in living tissue.
“Fibroblast migration plays a key role during various physiological and pathological processes such as wound healing and cancer metastasis,” said Bumsoo Han, a Purdue University professor of mechanical and biomedical engineering. “Although migration of individual fibroblasts has been well studied, migration in-vivo often involves simultaneous locomotion of groups of fibroblasts, so-called ‘en masse migration.”
New findings were published in October in the Journal of the Royal Society Interface.
“Specifically, we hypothesized that a group of migrating cells can significantly deform the matrix, whose mechanical microenvironment dramatically changes compared with the un-deformed state,” he said. “Because the cells cooperate, they can exert greater force in deforming the matrix compared to single cells alone. The alteration of the matrix microenvironment reciprocally affects cell migration.”
The hypothesis was tested by measuring the extracellular matrix deformation during en masse migration on collagen “hydrogels.” Findings showed the grouped cells act collectively to deform the matrix before and during the migration.
“We found that cells on soft collagen hydrogels migrate along tortuous paths, but, as the matrix stiffness increases, cell migration patterns become aligned with each other and show coordinated migration paths,” he said.
The research is related to work led by Andrew Mugler, a Purdue assistant professor of physics and astronomy, who co-authored a recent paper in the journal Physical Review Letters. The two researchers have collaborated for about two years.
“The common theme of both papers is how these cells cooperate,” Han said.
Findings will aid efforts to learn more about the en masse cell migration and its relationship to processes including cancer metastasis.
The Interface paper’s lead author is Purdue graduate student Altug Ozcelikkale. That paper probes how the cells “cooperate mechanically,” while the PRL paper focuses on how they cooperate chemically, Han said.
The research is ongoing, with future work investigating the multiple chemical and environmental cues involved in the collective cell behavior.
Writer: Emil Venere, 765-494-4709, venere@purdue.edu
Source: Bumsoo Han, 765-494-5626, bumsoo@purdue.edu
Cells that specialize in groups function more effectively
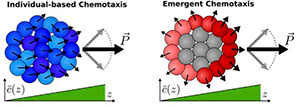 Cells track chemical gradients more precisely when they specialize (right), rather than acting as individuals (left).
Download image
Cells track chemical gradients more precisely when they specialize (right), rather than acting as individuals (left).
Download image
WEST LAFAYETTE, Ind. — Cells that specialize in groups track chemical gradients in their environment more precisely than individual cells, according to researchers at Purdue University.
Cells come together for collective migration in several processes, including organism development, wound healing and cancer metastasis. The simplest way for this to happen is by individual detection and response to a chemical attractant: each cell measures the difference in chemoattractant and moves in the direction of the gradient, while short-range coupling keeps the group together.
“Experiments suggest that in some cases, a new mechanism is emerging when cells get together as opposed to when they’re by themselves. We wanted to compare the accuracy of one of these emergent mechanisms to when cells act individually while part of a group,” said Andrew Mugler, an assistant professor of physics and astronomy at Purdue. “In particular, we wanted to know why they might have evolved this new mechanism – is there some advantage?”
There is. Cells that migrate as one-dimensional chains or two-dimensional sheets can detect gradients with a higher precision when they’re specializing in a group rather than stuck together as individuals, according to the study.
Cells can’t perfectly sense chemicals in their environment; there is always some error. When they come together in a group, they combine their sensory information, including errors. Individual cells working together in a group all contribute information and error, but in a group with the emergent mechanism, only the outer cells are sensory. This means there is less sensory information overall, but also less error, making that group better able to detect chemical gradients than their individually acting counterparts.
This emergent mechanism is also advantageous because if every cell in the group doesn’t need to perform every task, they exert less energy performing essential functions. Only the cells on the outskirts of the group would have to work to move it, for example.
Understanding how emergent cooperative sensing works will give researchers insight into several biological processes, including cancer metastasis, said Mugler. He plans to conduct further studies on collective cell behavior.
Writer: Kayla Zacharias, 765-494-9318, kzachar@purdue.edu
Source: Andrew Mugler, 765-496-3427, amugler@purdue.edu
New algorithm expands use of advanced camera for biological microscopy
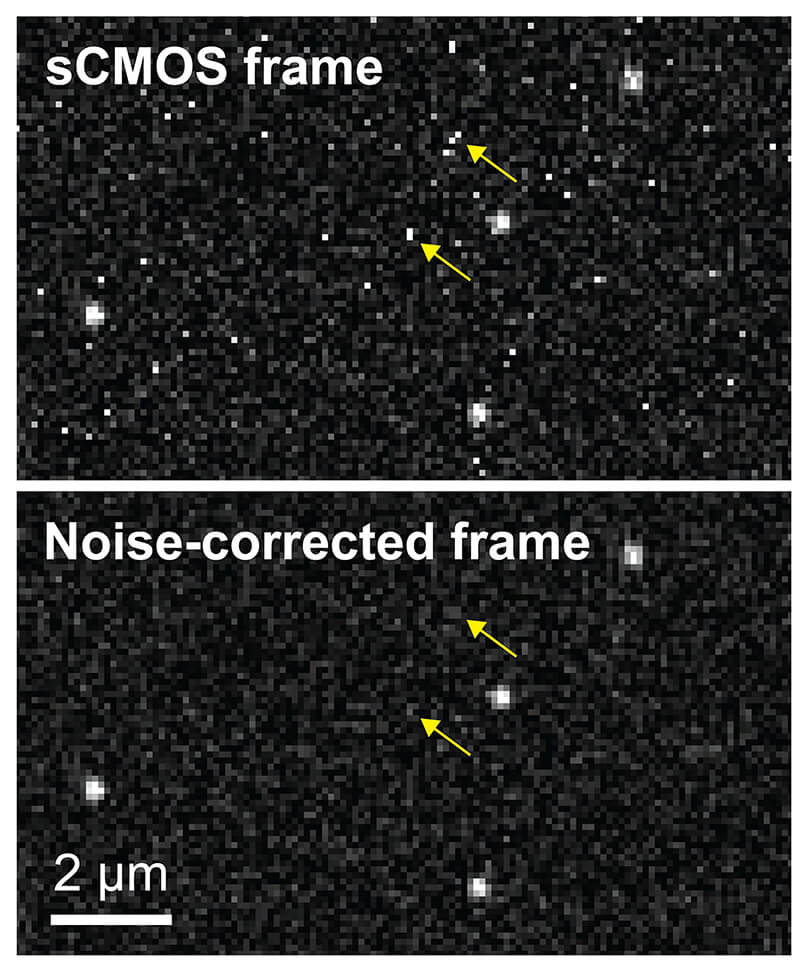 This photo shows the difference before and after use of the new algorithm. (Purdue University photo/ Sheng Liu and Fang Huang)
Download image
This photo shows the difference before and after use of the new algorithm. (Purdue University photo/ Sheng Liu and Fang Huang)
Download image
WEST LAFAYETTE, Ind. – A new computer algorithm allows scientists to use a high-performance sensor technology, called scientific complementary metal-oxide semiconductor cameras, for a wide range of biological research.
“Scientific sCMOS cameras are rapidly gaining popularity in biological sciences, material sciences and astronomy,” said Fang Huang, an assistant professor in Purdue University’s Weldon School of Biomedical Engineering. “The sensor provides significant advances in imaging speed, sensitivity and field of view compared with traditional detectors such as charge-coupled devices or electron multiplying CCD.”
However, the use of sCMOS cameras for biological research has been limited because of fluctuations in pixel quality, generating more “noise,” than the other cameras. In particular, every pixel fluctuates at its own rate.
“When you are trying to use this for biological studies, it's very difficult to determine whether this fluctuation comes from the sample (photons) or from the camera itself,” said Sheng Liu, the lead author of the paper, a postdoctoral research associate at the Weldon School of Biomedical Engineering.
Now, working with researchers in Purdue’s Department of Biological Sciences, Liu and Huang have developed a new algorithm that corrects the noise, making the sCMOS cameras available for a wide range of biological microscopy.
Findings were detailed in a research paper appearing earlier this year in the journal Nature Methods.
The authors include Liu; postdoctoral research associate Michael J. Mlodzianoski; graduate students Zhenhua Hu, Yuan Ren and Kristi McElmurry; Daniel M Suter, an associate professor in the Department of Biological Sciences; and Huang.
“We have been trying to use this camera for live-cell single-molecule super-resolution imaging and introduced an algorithm for that purpose in 2013,” Huang said. “However, the previous algorithm works only for single-molecule studies, which means all your objects have to be so-called point emitters. So, basically, your images must look like stars in the sky.”
Biological research, however, often involves imaging complex structures such as cellular organelles. To solve the problem, researchers developed the new algorithm.
“The fundamental challenge is to estimate one of the variables when you know the sum of two variables. There’s no unique answer to this question but we want to make the best estimate given our additional knowledge of the two variables.” Huang said. “We exploited a general property of imaging systems, the optical transfer function. Based on our knowledge of how each of the 4 million pixels on our camera chip behave, we are able to estimate the actual photon level at each pixel location. This is very exciting for us because this allows CMOS sensors to be used in a broad spectrum of imaging methods for quantitative biomedical and biological studies, improving their sensitivity, field of view and imaging speed”
The researchers filed a patent application on the algorithm through the Purdue Research Foundation’s Office of Technology Commercialization.
Portions of the research were funded by the National Institutes of Health (NIH (R35 GM119785)), the Defense Advanced Research Projects Agency (D16AP00093) and the National Science Foundation (1146944-IOS).
Writer: Emil Venere, 765-494-4709, venere@purdue.edu
Sources: Fang Huang, 765-494-6216, fanghuang@purdue.edu
Sheng Liu, 765-494-0863, liu1840@purdue.edu
