Study identifies enzymes that help fix cancer-causing DNA defects
April 22, 2014
 |
|
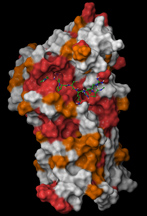
A model of the enzyme Cdc14. Red areas have identical amino acid sequences in human and yeast Cdc14. The ideal "key," or substrate, that would fit into Cdc14 is marked by the green and blue structure nestled into a "lock" that is the same in human and yeast Cdc14. (Molecular Cell / Mark Hall) |
WEST LAFAYETTE, Ind. - Purdue University researchers have identified an important enzyme pathway that helps prevent new cells from receiving too many or too few chromosomes, a condition that has been directly linked to cancer and other diseases.
Mark Hall, associate professor of biochemistry, found that near the end of cell division, the enzyme Cdc14 activates Yen1, an enzyme that ensures any breaks in DNA are fully repaired before the parent cell distributes copies of the genome to daughter cells. This process helps safeguard against some of the most devastating genome errors, including the loss of chromosomes or chromosome segments.
"It only takes one cell to start a tumor," Hall said. "This study gives us a platform for figuring out exactly what these enzymes are doing in human cells and how they impact genome stability and the avoidance of cancer."
Cdc14 has been linked to DNA damage repair in humans, but exactly how the enzyme helps preserve the genome and which proteins it regulates in this process have not been known.
 |
|
Mark Hall |
Hall and his fellow researchers developed a novel method of identifying the protein substrates upon which Cdc14 acts. Cdc14 regulates the function of other proteins by removing phosphate, a small chemical group, from them. Using Cdc14 in baker's yeast - which is very similar to human Cdc14 - the team studied the activity of the enzyme on a wide variety of synthetic substrate molecules, looking for similar features among the molecules most preferred by Cdc14.
"We were basically trying different keys in the lock to see which would fit the best," Hall said.
The team identified the most common structural features on molecules targeted by Cdc14 and used bioinformatics tools to pinpoint matching features in yeast proteins. Yen1 proved to be the best match, and further tests confirmed its role as a substrate of Cdc14. Yen1 is the first Cdc14 substrate involved in DNA repair to be identified.
Hall said the remarkable similarity of these enzymes in yeast and humans makes it likely that this method could be used to identify targets of Cdc14 in humans as well.
"Despite belonging to extremely different species, the 'lock' in yeast and human Cdc14 enzymes is exactly the same," he said. "That gives us confidence that we can use this strategy to identify substrates of human CDC14 and how they work to control DNA repair processes and prevent cancer."
Hall said understanding Cdc14's role in DNA repair and how the enzyme binds to its substrates could be used to develop more effective chemotherapeutic weapons against cancer. Many chemotherapeutic drugs work by producing such extensive DNA damage in cancer cells that they kill themselves. Designing a chemical that mimics the features of a Cdc14 substrate would help block Cdc14 from repairing damaged DNA in cancer cells, speeding their death.
"Developing Cdc14 inhibitory compounds could make certain cancer treatments more specific and potent," Hall said. "You could think of Cdc14 inhibitors as kryptonite to cancer cells, potentially weakening their ability to heal themselves and making them more vulnerable to chemotherapy treatment."
Hall also is exploring the possibility of using Cdc14 inhibitors to combat deadly fungal diseases in crops.
The paper was published in Molecular Cell. A summary is available at http://www.cell.com/molecular-cell/abstract/S1097-2765%2814%2900128-2
Collaborators on the research include Purdue graduate students Christie Eissler and Brendan Powers, Lorraine Symington and Gerard Mazón of the Columbia University Medical Center and Sergey Savinov of the Purdue University Center for Cancer Research.
Funding for the research was provided by the National Institutes of Health and the Purdue University Center for Cancer Research.
Writer: Natalie van Hoose, 765-496-2050, nvanhoos@purdue.edu
Source: Mark Hall, 765-494-0714, mchall@purdue.edu
ABSTRACT
The Cdk/Cdc14 module controls activation of the Yen1 Holliday junction resolvase to promote genome stability
Christie L. Eissler 1, 2; Gerard Mazón 3; Brendan L. Powers 1, 2; Sergey N. Savinov 2, Lorraine S. Symington 3; Mark C. Hall 1, 2
1 Department of Biochemistry, Purdue University, West Lafayette, IN 47907, USA
2 Center for Cancer Research, Purdue University, West Lafayette, IN 47907, USA
3 Department of Microbiology and Immunology, Columbia University Medical Center, New York, NY 10032, USA
E-mail: mchall@purdue.edu
Faithful genome transmission during cell division requires precise, coordinated action of DNA metabolic enzymes, including proteins responsible for DNA damage detection and repair. Dynamic phosphorylation plays an important role in controlling repair enzymes during the DNA damage response (DDR). Cdc14 phosphatases oppose cyclin-dependent kinase (Cdk) phosphorylation and have been implicated in the DDR in several model systems. Using new insight into Cdc14 specificity we identified the budding yeast Holliday junction resolvase Yen1 as the first DNA repair target of Cdc14. Cdc14 activation at anaphase triggers nuclear accumulation and enzymatic activation of Yen1, likely to resolve persistent recombinational repair intermediates. Consistent with this, expression of a phosphomimetic Yen1 mutant increased sister chromatid non-disjunction. In contrast, lack of Cdc phosphorylation resulted in constitutive activity and elevated crossover-associated repair. The precise timing of Yen1 activation governed by core cell cycle regulators helps coordinate DNA repair with chromosome segregation and safeguards against genome destabilization.
Ag Communications: (765) 494-2722;
Keith Robinson, robins89@purdue.edu
Agriculture News Page

