
July 12, 2005
Biologists see combined structure of cold virus and receptor molecule
WEST LAFAYETTE, Ind. – Biologists at Purdue University have determined the combined structure of a common-cold virus attached to a molecule that enables the virus to infect its host, information that ultimately may help researchers develop methods for treating certain viral infections.
|
Coxsackievirus A21 infects host cells first by recognizing a "receptor molecule" called ICAM-1, which is located on the cell's surface, and then by anchoring itself to the molecule. ICAM-1 stands for intracellular adhesion molecule 1.
"ICAM-1 is the same receptor molecule used by the vast majority of viruses that cause the common cold," said Chuan Xiao, a doctoral student who is leading the research in the laboratory of Michael Rossmann, the Hanley Distinguished Professor of Biological Sciences in Purdue's College of Science.
Findings will appear in the July issue of the journal Structure.
"The real objective of this work is to study the whole complex of ICAM-1 and the virus as a single entity," Rossmann said. "Being able to characterize the combined structure of the virus and ICAM-1 will teach us how the virus recognizes a particular kind of molecule and how it then anchors to the cell, which represents the initial stages of infection."
Ultimately, researchers are trying to learn more about the binding mechanisms because such knowledge might eventually lead to new treatments.
"One of the many different ways of inhibiting viral infection is to stop the virus from binding to cells," Rossmann said. "That has not been our objective in this case. We just want to learn how this virus infects its host cell. In other words, how does the virus get into the host?"
Coxsackievirus A21 is one of several viruses that cause the common cold.
The researchers used two methods to learn the structure of the virus-molecule complex. One method, a technique called X-ray crystallography, yielded images of the virus with a resolution of 2.5 angstroms, which is nearly fine enough to see individual atoms. Using this technique, researchers create crystals of a substance, in this case the virus. Then, X-rays are passed through the crystals, creating a "diffraction pattern" that can be interpreted with various computational procedures to produce an image.
The other method, a powerful imaging tool called cryo-electron microscopy, was used to determine the entire three-dimensional structure of the virus-molecule complex. With this technique, specimens are first frozen before they are studied with an electron microscope. Cryo-electron microscopy enables scientists to study details as small as 8 angstroms, resolution high enough to see groups of atoms. An angstrom is one ten-billionth of a meter, or roughly a millionth as wide as a human hair.
"The electron microscopy is necessary for studying the entire complex because you can't crystallize the complex of ICAM-1 and the virus," Rossmann said. "That's because crystallization often takes days, weeks or months, but the complex is only stable for hours, which means it doesn't stay together long enough to crystallize."
The researchers pieced together the overall structure of the virus and ICAM-1 by combining the high-resolution X-ray crystallography images of the virus with the lower resolution electron microscope view.
The findings represent the first time researchers have seen fine details of the complex's structure.
"It's important to see the shape of the complex because that could tell us how the virus recognizes the host cell," Rossmann said. "Knowing the structure might also reveal the initial stages of what happens after attachment, and indeed there probably are different steps in the attachment process.
"The receptor apparently binds into what we call a canyon, which is a surface depression on the virus, and it might do that in at least two different steps. Perhaps it binds once loosely on the surface, and then it might bind again deeply into the canyon to strengthen its attachment."
The research paper was written by Xiao; Carol M. Bator-Kelly, a technical assistant in Rossmann's lab; Elizabeth Rieder, a technical assistant for Eckard Wimmer, a virologist at the State University of New York at Stony Brook; Paul R. Chipman, an electron microscopist at Purdue; Alister Craig, a researcher from the Liverpool School of Tropical Medicine in the United Kingdom; Richard J. Kuhn, a professor of biological sciences at Purdue; Wimmer; and Rossmann.
The images are revealing new details about how amino acids, which are the building blocks of proteins, interact during the binding of Coxsackievirus A21 and ICAM-1.
"In general, some amino acids have a positive charge, and some have a negative charge, and these opposite charges can play a critical role in attracting a virus to a host cell," Rossmann said. "It turns out the attraction between negative and positive charges on the virus and on the host cell seem to be a dominating feature but not the entire story of the recognition process.
"There is also shape involved, and the canyon on the virus and features on the ICAM-1 molecule have to match each other, like a key going into a keyhole."
The research has been funded primarily through a grant from the National Institutes of Health.
Xiao said the research is ongoing, and future work may delve into how Coxsackievirus A21 binds to another cell-receptor molecule called DAF, or decay acceleration factor. The virus binds to both ICAM-1 and DAF at the same time, so future work may result in finding the structure of a complex that includes both molecules attached to the virus.
Writer: Emil Venere, (765) 494-4709, venere@purdue.edu
Sources: Michael Rossmann, (765) 494-4911, mgr@indiana.bio.purdue.edu
Chuan Xiao, xiaoc@purdue.edu
Purdue News Service: (765) 494-2096; purduenews@purdue.edu
Note to Journalists: Copies of the research paper are available by calling the press office of the journal Structure at (617) 397-2879, or through EurekAlert!. A publication-quality image showing the structure of the virus-molecule complex is available from Emil Venere, (765) 494-4709, venere@purdue.edu, and it can be downloaded .
IMAGE CAPTION:
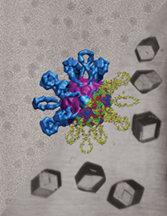
This composite image shows the combined structure of Coxsackievirus A21 and a "receptor molecule" called ICAM-1, or intracellular adhesion molecule 1. The virus is one of the viruses that causes the common cold, and the receptor molecule enables the virus to attach to and infect host cells. ICAM-1, located on the surfaces of cells, is represented in blue, and the virus is represented as red. Researchers at Purdue University have determined the structure of the virus-molecule complex by combining images taken using X-ray crystallography and cryo-electron microscopy. Being able to determine the combined structure of the virus and ICAM-1 may teach scientists how the virus recognizes the molecule and how it then anchors to the cell, which represents the initial stages of infection. The virus-molecule complex in the center of this image is a thousandth as wide as a human hair. (Graphic/Department of Biological Sciences, Purdue University)
A publication-quality graphic is available at https://www.purdue.edu/uns/images/+2005/rossmann-coldvirus.jpg
SUMMARY
The Crystal Structure of Coxsackievirus A21
and its Interaction with ICAM-1
Chuan Xiao1, Carol M. Bator-Kelly1, Elizabeth Rieder 2, Paul R. Chipman1, Alister Craig3, Richard J. Kuhn1, Eckard Wimmer2, and Michael G. Rossmann1.
1
Department of Biological Sciences, Purdue University2
Department of Molecular Genetics and Microbiology, School of Medicine, State University of New York at Stony Brook3
Liverpool School of Tropical MedicineCVA21 and poliovirus belong to the Enterovirus genus in the family of Picornaviridae, whereas rhinoviruses form a distinct picornavirus genus. Nevertheless, CVA21 and the major group of human rhinoviruses recognize intercellular adhesion molecule 1 (ICAM-1) as their cellular receptor, whereas polioviruses use poliovirus receptor. The crystal structure of CVA21 has been determined to 3.2-angstrom resolution. Its structure has greater similarity to poliovirus structures than to other known picornavirus structures. Cryo-electron microscopy (cryo-EM) was used to determine an 8.0-angstrom resolution structure of CVA21 complexed with an ICAM-1 variant, ICAM-1Kilifi. The cryo-EM map was fitted with the crystal structures of ICAM-1 and CVA21. Significant differences in the structures of CVA21 with respect to the pilogivur structures account for the inability of ICAM-1 to bind polioviruses. The interface between CVA21 and ICAM-1 has shape and electrostatic complementarity with many residues being conserved among those CVAs that bind ICAM-1.
To the News Service home page