 Purdue News
Purdue News
 Purdue News
Purdue News

|
|
March 2000 Virus study reveals how nature
|
|
"It appears that these viruses don't come together all at once but are assembled in pieces, like an automobile, with small subunits coming together to form larger components of the structure," says Timothy S. Baker, professor of biological sciences at Purdue and principal investigator on the study.
The use of small three-sided clusters to make larger structures may be a common tactic used by large viruses to develop stable and highly symmetrical structures, Baker says, noting that many virus shells are made from a combination of five- and six-sided clusters.
Knowing the structure of these large viruses may provide insights on how other large viruses are assembled, and may someday help scientists find ways to design antiviral agents, he says.
The viruses – Chilo iridescent virus, a virus that infects the rice stem borer insect, and Paramecium bursaria chlorella virus type 1, a virus that infects certain freshwater algae – are among the largest known viruses. They are about 200 times more massive than the common cold virus.

|
"These viruses contain well over 5,000 protein subunits," Baker says. "To form a structure of that size, so perfectly and with such symmetry, raises some mind-boggling questions about how something so precise can form from all of these individual molecules."
The process also raises some complex geometry and terminology.
Though the viruses look similar to their smaller counterparts – many of which contain as few as 180 protein subunits – the outer shells of the super-size viruses contain thousands more components.
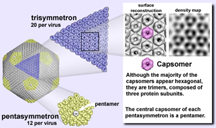
Each shell, or capsid, is a quilt-work pieced together from more than 5,000 individual protein subunits. These subunits join to form two types of doughnut-shaped clusters, or capsomers. One capsomer contains three protein subunits, called a trimer, and the other is composed of five protein subunits, called a pentamer. The virus shell consists of 1,680 trimers and 12 pentamers.
The trimers and pentamers link to create large triangular and pentagonal building blocks – called trisymmetrons and pentasymmetrons, respectively.
"These large viruses are distinct because they have large numbers of these trimers that come together to form larger structures on the virus shell," Baker says. "It is the symmetry created when all these individual molecules come together that gives the virus shell its strength and makes it stable."
Two smaller viruses, the adenovirus and the bacteriophage PRD 1 virus – also have shells that contain trimers, indicating that the viruses may have an evolutionary link to the Chilo and the Paramecium viruses.
The study also shows that the large viruses each contain a lipid bilayer membrane within their shell, a feature that is somewhat unusual in viruses, Baker says. The study shows how the membrane is organized and structured.
The sheer size of the viruses – due to the fact that they carry a lot of their own "machinery" needed to infect a cell – also sets them apart from other viruses.
"Most simpler viruses, like rhinovirus, carry a genome that allows them to go into a cell and take over the host-cell's machinery in order to replicate," Baker says. "These viruses, in addition to their genome, carry all the enzymes needed to dissolve the host cell wall, deposit their genomes and carry out new rounds of DNA replication."
This presence of many cellular components, together with the common structural features found in the study, suggests that the two viruses may have evolved from a single-celled organism, Baker says.
"These may be very ancient viruses that somehow got caught in the early stages of evolving," he says. "While many viruses evolved and acquired the ability to let the cell do everything for them, these viruses may have just lumbered along, carrying all of this machinery with them."
Some single-celled organisms contain just slightly more genetic material than these viruses, he adds.
The virus studies were done at Purdue using the techniques of cryo-electron microscopy and three-dimensional image reconstruction. The techniques, which use hundreds of two-dimensional images to develop a three-dimensional view of the virus, allowed the group to determine the overall shape of virus particles and see the symmetrical arrangement of their components.
"These techniques have helped revolutionize the study of virus assembly, especially for viruses too large to study with traditional techniques such as X-ray crystallography," Baker says. "We actually had some of our first images of these viruses nine years ago, but we couldn't process them because we didn't have computers that could handle the tremendous amount of data."
The Purdue group now is using the techniques to study the African swine fever virus, one of the largest known icosahedral viruses.
The Purdue team included graduate research assistant Xiaodong Yan; Norman Olson, senior research electron microscopist; and Michael Rossmann, the Hanley Distinguished Professor of Biological Sciences. In addition, the Purdue group collaborated with James Van Etten of the University of Nebraska and Max Bergoin of the Laboratory of Comparative Pathology in Montpellier, France.
The studies were funded by the National Institutes of Health, the National Science Foundation and the Lucille P. Markey Foundation.
Source: Timothy Baker, (765) 494-5645, tsb@bragg.bio.purdue.edu
Writer: Susan Gaidos, (765) 494-2081; sgaidos@purdue.edu
Purdue News Service: (765) 494-2096; purduenews@purdue.edu
Related Web site:
Structural studies at Purdue
ADDITIONAL INFORMATION:
An interactive animation of the virus structure, this screen shot from the animation, and a publication-quality photo of Timothy S. Baker are available at the News Service Web site and at the ftp site. (Purdue graphic by Clark Gedney).
 To the Purdue News and Photos Page
To the Purdue News and Photos Page